Fighting viruses with mathematical modelling
Posted on 6 January 2015
We've produced a mathematical model that shows how viruses behave and could help in the development of anti-viral therapies.
This truly interdisciplinary effort has provided surprising insights into a fundamental mechanism in virology.”
The model, which was constructed in partnership with the University of Leeds, provides important new insights about the molecular mechanisms behind virus assembly.
Anti-viral therapies
The discovery opens up new possibilities for the development of anti-viral therapies and could help in the treatment of a range of diseases from HIV and Hepatitis B and C to the “winter vomiting bug” Norovirus and the Common Cold.
The researchers, led by Professor Reidun Twarock from our Departments of Mathematics and Biology, have established a theoretical basis for the speed and efficiency with which viruses assemble protective protein containers for their genetic information – in this case an RNA molecule – during an infection.
By incorporating multiple specific contacts between the genomic RNA and the proteins in the containers, and other details of real virus infections, the research team’s mathematical model demonstrates how these contacts act collectively to reduce the complexity of virus formation.
Interdisciplinary approach
Professor Twarock, a member of the York Centre for Complex Systems Analysis (YCCSA), said: “This truly interdisciplinary effort has provided surprising insights into a fundamental mechanism in virology. Existing experimental techniques for studying viral assembly are unable to identify the co-operative roles played by all the important components, highlighting the need and power of mathematical modelling. This model is a paradigm shift in the field of viral assembly. It sheds new light on virus assembly in a major class of viruses and their evolution, and opens up a novel strategy for antiviral therapy.”
The research, published in the Proceedings of the National Academy of Sciences (PNAS), was funded by the Engineering and Physical Sciences Research Council, the Biotechnology and Biological Sciences Research Council and the University of York.
The text of this article is licensed under a Creative Commons Licence. You're free to republish it, as long as you link back to this page and credit us.

Professor Reidun Twarock
Her research focuses on the development of mathematical and computational tools to model fundamental processes in virology
Discover the details
Find out more in the York Research Database
Article
- Solving a Levinthal's paradox for virus assembly identifies a unique antiviral strategy (Proceedings of the National Academy of Sciences)
Visit the departments
Explore more research
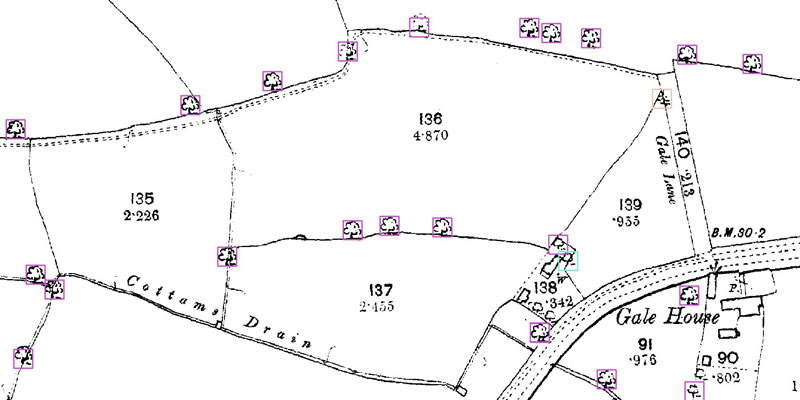
A research project needed to spot trees on historic ordnance survey maps, so colleagues in computer science found a solution.

We’re using gaming technology to ensure prospective teachers are fully prepared for their careers.

A low cost, high-accuracy device, could play a large part in the NHS's 'virtual wards'.
Research newsletter
Our monthly research newsletter features a curated mix of news, events, and recent discoveries delivered straight to your inbox.
