News

The Green Chemistry Centre has achieved and been awarded My Green Lab® Certification for Green Chemistry York at the Platinum Level.
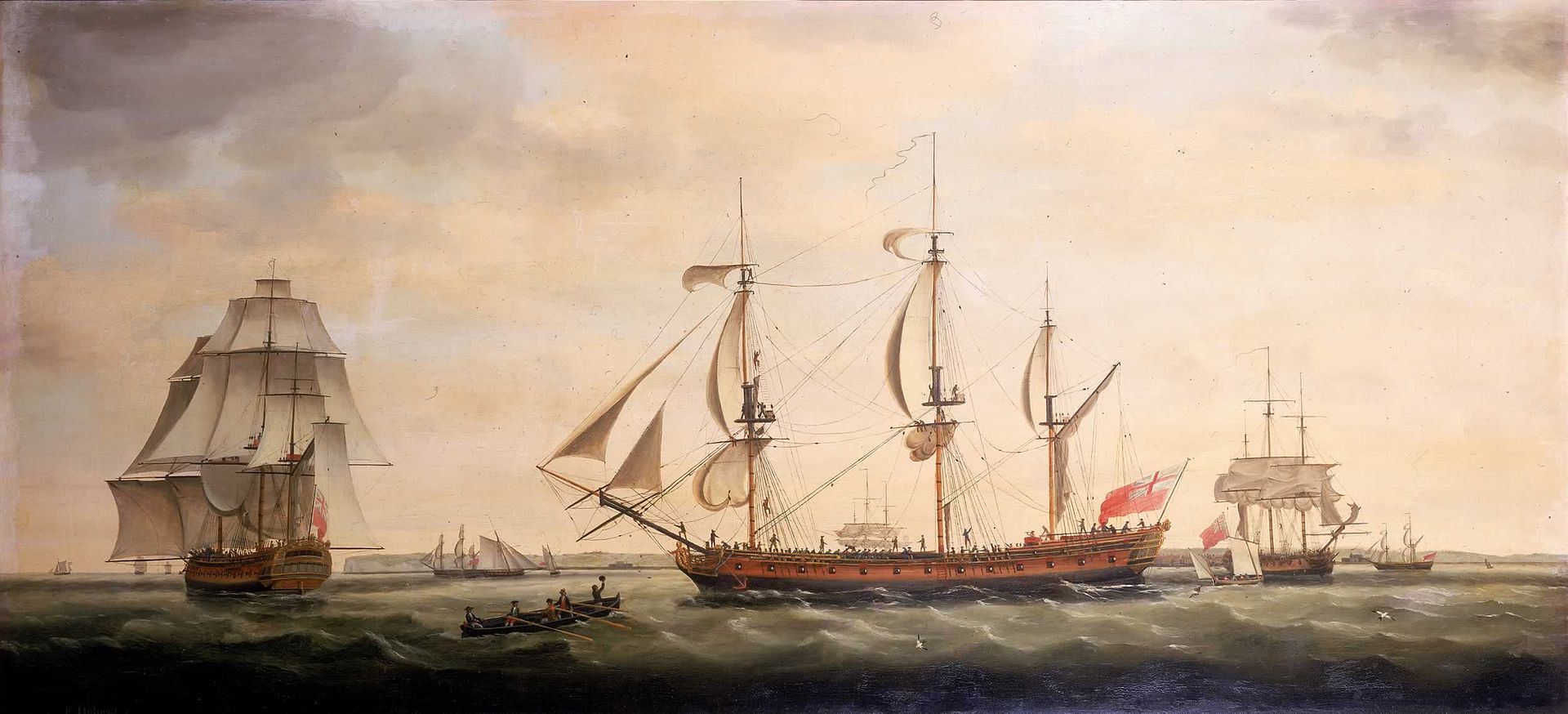

A new record of global warming pushes our understanding back to the very start of the industrial revolution using old weather reports from sailing ships.

The TransPharm consortium have collaborated to create an educational website explaining the environmental impact of pharmaceuticals and describing safer and more sustainable practices in the healthcare sector. The platform can be found at www.sustainablepharmaceuticals.eu.
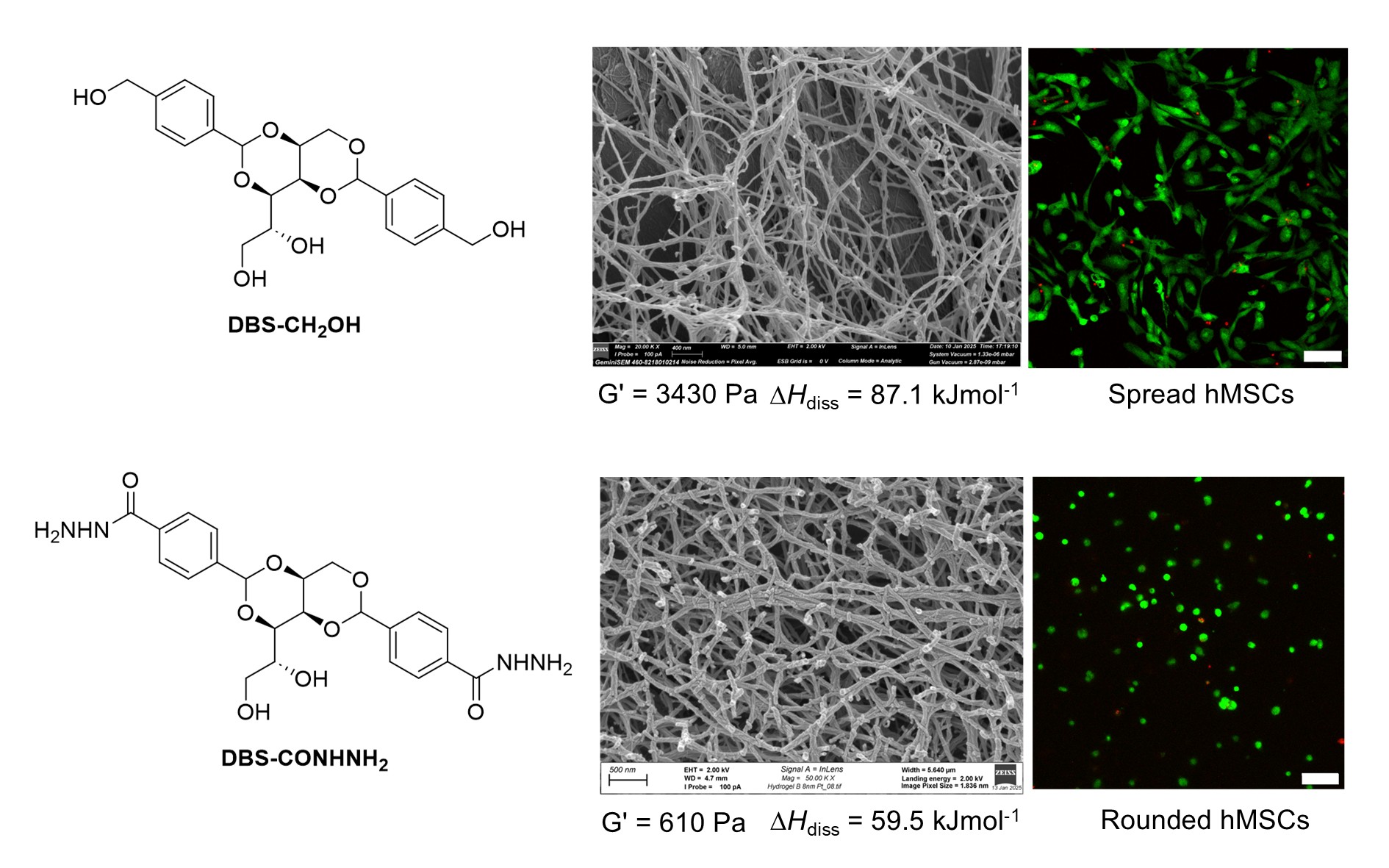
Scientists from the University of York have developed innovative self-assembling gels that direct and control the growth of human stem cells, with potential applications in regenerative medicine.

Alfonso Burri Mereles, a Chevening Scholar from Paraguay, explains how he overcame setbacks to pursue his passion for Green Chemistry at the University of York.

Professor Paul Walton has been elected as a foreign member of the Royal Swedish Academy of Sciences (RSAS).

Two new doctoral training awards at the University of York will be used to train the next generation of scientists through specialised PhD programmes. They will equip these new researchers with the technical and transferable skills needed to contribute to the UK’s bioeconomy, while fostering a collaborative and inclusive training environment.
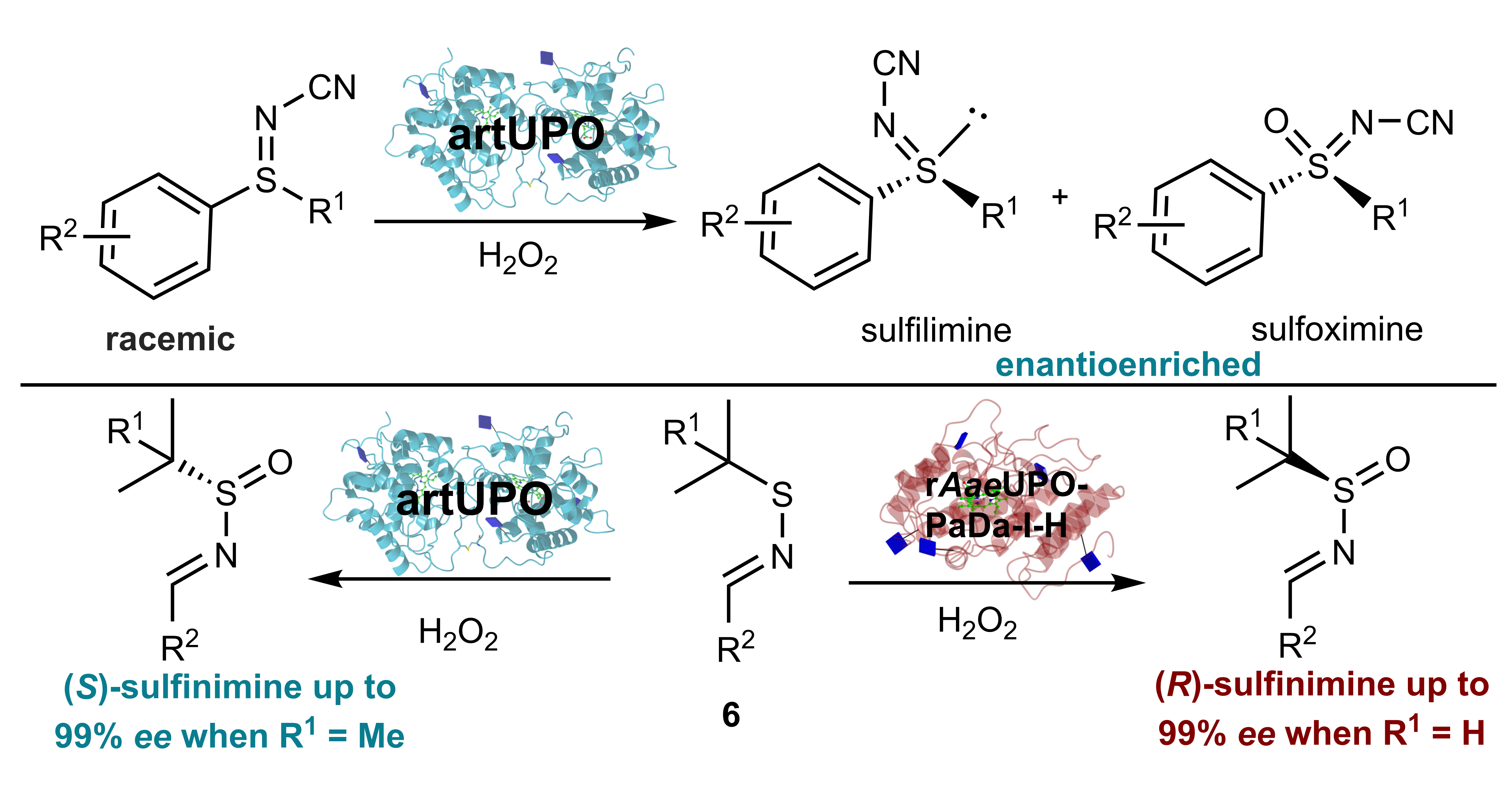
A new method for the preparation of medicinally important chiral sulfur compounds has been developed using a class of enzymes called ‘Unspecific Peroxygenases’.
ChemYork Magazine
We produce a biannual magazine highlighting our teaching, research, staff and students.
Read ChemYork
