Cell and Developmental Biology
Research highlights
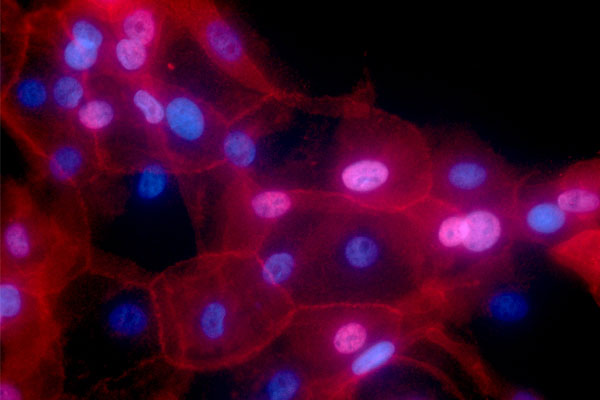
Electrical charges advancing spread of breast cancer
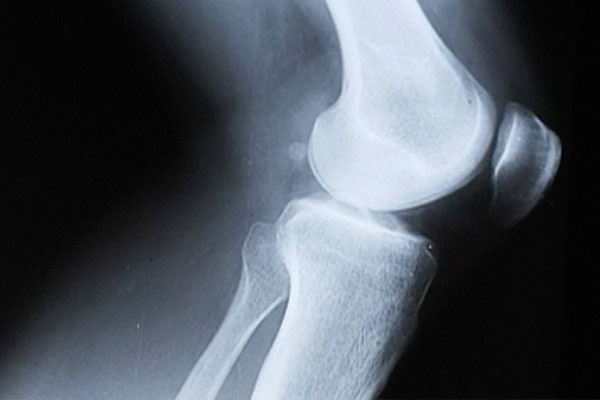
Altering sugar chains to promote bone growth
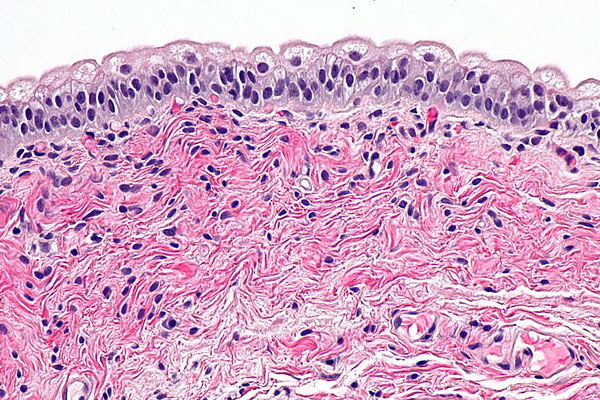
Moving closer to solving chronic bladder diseases
People
| Name | Expertise |
| Nia Bryant |
|
| William Brackenbury |
|
| Sangeeta Chawla |
|
| Dawn Coverley |
|
| Paul Genever |
|
|
Darren L Goffin |
|
| Harry Isaacs |
|
| Chris MacDonald |
|
| Betsy Pownall |
|
| Paul Pryor |
|
| Nathalie Signoret |
|
| Jenny Southgate |
|
| Sean Sweeney |
|
| Daniel Ungar |
|

