Professor Daniel Ungar
Professor
Research
Overview
The diversity of eukaryotic glycan structures is produced by a large number of Golgi localized glycosylation enzymes. The precise localization of these enzymes to different Golgi cisternae is essential for accurate glycan synthesis. We would like to understand how the organization of the glycosylation machinery governs glycan biosynthesis.
- More of our research can be found on the Ungar Lab website
Vesicle tethering at the Golgi and glycosylation
The conserved oligomeric Golgi (COG) tethering complex is responsible for coordinating vesicle targeting to sort glycosylation enzymes to their respective cisternae. We have previously mapped protein-protein interactions between the eight mammalian COG subunits and members of other trafficking protein families. More recent projects have investigated the (mis)localization of enzymes within the Golgi upon COG-mutations, and the resulting changes in glycosylation. We are also generating mutations in COG subunits that alter COG’s interaction with selected binding partners with the aim to obstruct a subset of the intra-Golgi vesicles. The characterization of these mutants is done with the support of biologics manufacturers interested in new ways for glyco-engineering mammalian cells.
Glycan profiling and modelling glycan biosynthesis
To understand the consequences of altered enzyme sorting within the Golgi we are developing analytical tools to profile the glycans made by the Golgi. The filter-aided N-glycan separation method was originally developed for N-glycans, and then expanded for O-glycans as well. A further addition to our toolkit is a stochastic computational model of N-glycan processing in the Golgi. By using Bayesian fitting between modelled and experimentally determined glycan profiles, we can generate predictions about the changes of Golgi organization upon COG mutations. The profiling and modelling tools are now employed in projects with strong industrial links to investigate glycosylated biologics and the (bio)synthesis of human milk oligosaccharides.
Investigating glycan functions during cellular differentiation
To test the concept of “forward glycomics”, we started using a mesenchymal stem cell model that can differentiate into osteoblasts. Changing glycosylation using COG-defects could indeed alter differentiation. Interestingly, we found enhanced osteogenic differentiation caused by inhibiting complex N-glycan formation, something that our modelling suggests could be due to a shift in glycan branching during osteogenesis.
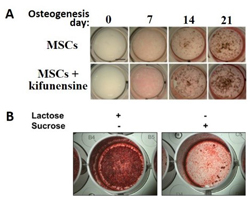
Figure 1. Glycans influence osteogenic differentiation of MSCs.
Three week differentiation of MSCs ± the glycan processing inhibitor kifunensine (A) or in the presence of the sugars lactose and sucrose (B). Mineralization was stained by von Kossa staining (brown specs – A) or alizarin red staining (red colour – B).
Teaching and scholarship
![]()
I believe that different students need different teaching styles, yet each student needs a chance to maximise their potential. Therefore, while lecturing may be the most efficient way to deliver information to large numbers of students, I put a lot of emphasis on teaching in smaller groups, such as tutorials, final year projects and practicals, where students’ different learning styles can be much more readily accommodated, and therefore their full potential can often be better achieved.
![]()
Lectures are of course still very important, and I make use of my research again to teach diverse areas of molecular cell biology. To make it more accessible and interesting to the large groups I focus on teaching concepts rather than factual details. Wherever I can I also include some glycobiology in my teaching. This is an area of research strength in York, so I feel it is important to prepare students for independent projects that may focus on this interesting field.
![]()
Tutorials are especially versatile, and we generally talk about topics on organelles and glycobiology. It is easy to excite students while teaching them useful skills and scientific concepts in these sessions. Students enjoy these very much, and are ultimately happy with the set of skills acquired.
![]()
I particularly enjoy supervising final year projects students. By providing both dry and wet-lab based projects centred around my research interests of molecular cell biology and glycobiology, the students are integrated into a vibrant research group of PhD students and postdoctoral fellows. For most students this is their first experience with an independent research project, and it is a joy to see their development into critical thinkers and industrious laboratory workers.


Contact details
https://sites.google.com/york.ac.uk/ungar-lab/research
