Dr Michael Plevin
Senior Lecturer
Research
My research addresses mechanisms of biomolecular recognition and the structural and chemical features that define interaction surfaces of proteins and nucleic acids. Within this broad subject area, I focus on multi-domain proteins involved in regulation and signalling in humans and human diseases, and how the interplay between structured domains and intrinsically disordered regions relates to function and mis-function in modular proteins.
My research group uses a variety of biochemical, biophysical and structural techniques to explore the sequence-structure-function relationship of proteins. We work in close collaboration with molecular cell biologists at York and other national and international research centres, as well as with scientists in industry.
A major research focus is the structural biophysics of protein and RNA molecules involved in RNA silencing and human microRNA (miRNA) biogenesis. In recent years, this has concentrated on how Dicer co-factors interact with and recognize miRNA precursors, and how the mechanisms underpinning their association has been conserved for over 500 million years, from insects to humans. This ongoing work is the subject of several recent and current BBSRC- and WT-funded projects, including a 2016 BBSRC New Investigator award.
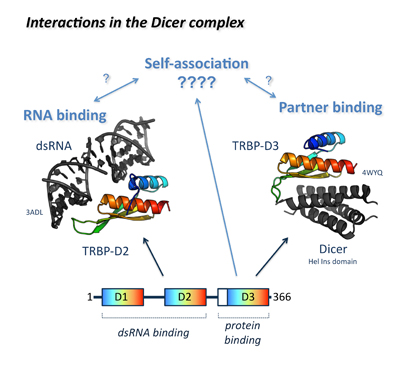
Figure 1. Interplay between inter- and intramolecular interactions in the assembly and function of the Dicer complex.
A second theme relates to non-covalent interactions involving aromatic groups in biomacromolecules. We have developed approaches for detecting aromatic hydrogen-bond-like interactions in proteins and determined their contribution to protein-protein recognition. These studies provided the first definitive experimental evidence for the existence of two types of XH/pi interaction in proteins. Our current work investigates how these weak interactions contribute to the structure, dynamics and function of proteins.
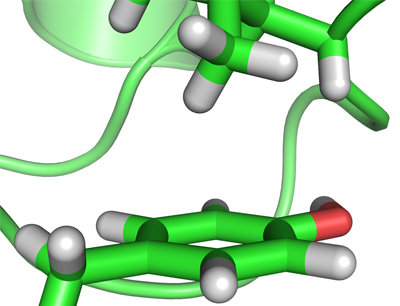
Figure 2. XH/pi interactions in proteins: Tyrosine-59 and Leucine-50 in human ubiquitin (1UBQ)
Liquid-liquid phase separation (LLPS) is a fascinating area of biophysical science that sits nicely at the interface of our interests in non-covalent interactions, multi-domain proteins and RNA. My group has a developing interest in this phenomenon, particularly the role of modular proteins, intrinsic disorder and weak transient interactions in the process of phase separation, how phase separation underpins the emergence of organization and function in the cell, and the use of solution biophysical techniques to characterize these processes.
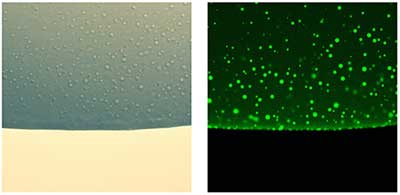

Figure 3. Studying biomolecular phase separation in vitro using recombinant proteins and light and fluorescence microscopy.
Lastly, my group has ongoing collaborations with industrial partners in which we seek to exploit the properties of natural proteins for applications in biotechnology. Current collaborations include Oxford Nanopore Technologies (Oxford, UK), Aptamer Solutions (York, UK) and NMR-Bio (Grenoble, France).
Our work has been funded by organisations such as BBSRC, MRC, The Wellcome Trust and The Royal Society.
Teaching and scholarship
![]()
My teaching portfolio at York is heavily influenced by the activities of my research group, the biological processes we study, and the techniques we use.
![]()
My taught material falls into two main categories. I deliver fundamental training in the skills and techniques needed for modern biochemical research. In Stage 2, I contribute to courses that introduce the principles and applications of biochemical research approaches, including techniques to determine and characterize the 3D structures of biomacromolecules (proteins, nucleic acids, etc) and to describe how molecules interact (how tightly, how specifically, for how long, etc). I run practical laboratory training in Stages 2 and 3 that develops and applies techniques such as purifying recombinant proteins, quantifying protein activity, and evaluating protein structure-function relationships via site-directed mutagenesis. In Stage 3, I contribute to lecture-based modules that explore molecular recognition in important biological contexts. My lectures address how proteins recognize sequence and structural features in protein and RNA molecules; how multi-domain proteins combine different modules to deliver function; how both structure and unstructured regions of proteins are critical for function. In each case, these phenomena are explored through core cellular processes and linked to experimental approaches and data reported in the literature.
![]()
The Stage 3 and 4 projects available in my research group align closely with our research activities. Themes of protein modularity, multi-protein complexes, macromolecular recognition, sequence-structure-function relationships will be central to any project we supervise. We aim to train project students in advanced laboratory techniques, including thermodynamic analyses of protein/protein interactions, NMR spectroscopy and protein crystallography as well as computational analyses of protein structures.


