Dr Seishi Shimizu
Reader in Biochemistry
Overview
Statistical Thermodynamics: Theory and Applications
I am a theoretician who aims:
- to gain a molecular scale understanding directly from the routinely performed experiments (such as solubility, sorption isotherm and physical properties' data);
- to attain interpretive clarity via a direct connection to the first principles with a bare minimum number of assumptions;
- to make my theory usable for industry and experiments via user-friendly, web-based apps (written by my industrial collaborator).
We work in the following research areas.
Solution phenomena
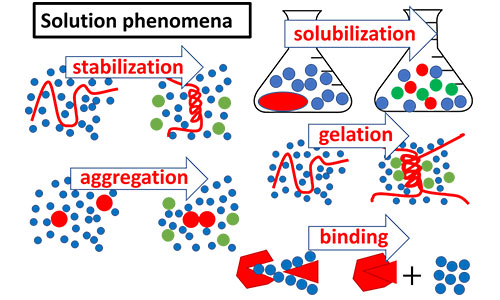
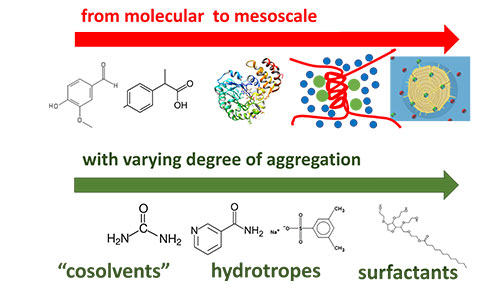
Interfacial phenomena
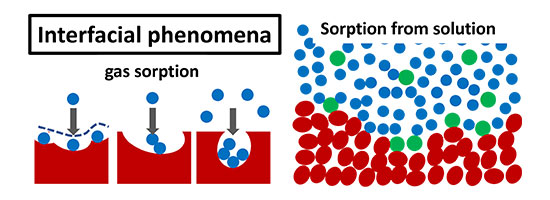
I strive for clarity using pen, paper, coffee and walks. I collaborate with theoreticians, experimentalists and industry.
To find out more:
- Cooperativity in Sorption Isotherms: Shimizu and Matubayasi, Langmuir 39, 2023, 13820-13829.
- Understanding Sorption Mechanisms Directly from Isotherms: Shimizu and Matubayasi, Langmuir 39, 2023, 6113-6125.
- Surface Area Estimation: Replacing the Brunauer–Emmett–Teller Model with the Statistical Thermodynamic Fluctuation Theory: Shimizu and Matubayasi, Langmuir 38, 2022, 7989-8002.
- Formulating rationally via statistical thermodynamics: Shimizu, Curr. Opin. Coll. Interf. Sci. 2020, 48, 53-64.
- Unifying hydrotropy under Gibbs phase rule: Shimizu & Matubayasi, Phys. Chem. Chem. Phys. 2017, 19, 23597.
- Practical molecular thermodynamics for greener solution chemistry: Abbott, Booth and Shimizu, Green Chem. 2017, 19, 68-75.
- “Gastrophysics” Shimizu, Stenner and Matubayasi, Food Hydrocoll. 2017, 62, 128-139.
- Food flavour: Shimizu, Abbott and Matubayasi, Food & Function, 2017, 8, 2999-3009.
Publications
Publications
My publications
For an up-to-date full list of my publications, please see the York Research database.
Selected publications
- Cooperativity in Sorption Isotherms
S Shimizu and N Matubayasi, Langmuir 2023, 39, 13820-13829. - Understanding Sorption Mechanisms Directly from Isotherms
S Shimizu and N Matubayasi, Langmuir 2023, 39, 6113-6125. - Sorption from Solution: A Statistical Thermodynamic Fluctuation Theory
S Shimizu and N Matubayasi, Langmuir, 2023, 39, 12987-12998. - Surface Area Estimation: Replacing the Brunauer-Emmett-Teller Model with the Statistical Thermodynamic Fluctuation Theory
S Shimizu and N Matubayasi, Langmuir, 2022, 585, 7989-8002. - Ensemble transformation in the fluctuation theory
S Shimizu and N Matubayasi, Physica A, 2022, 585, 126430 - Cooperative sorption on porous materials
S Shimizu and N Matubayasi, Langmuir, 2021, 37, 10279–10290 - Sorption: a statistical thermodynamic fluctuation theory
S Shimizu and N Matubayasi, Langmuir, 2021, 37, 7380–7391 - Temperature dependence of sorption
S Shimizu and N Matubayasi, Langmuir, 2021, 37, 11008–11017 - Thermodynamic stability condition can judge whether a nanoparticle dispersion can be considered a solution in a single phase
S Shimizu and N Matubayasi, J Coll Interf Sci.,2020, 575, 472-479 - Formulating rationally via statistical thermodynamics
S Shimizu, Curr Opin Coll Interf Sci, 2020, 48, 53-64. - Water Networks Can Determine the Affinity of Ligand Binding to Proteins
Darby JF, Hopkins AP, Shimizu S, Roberts SM, Brannigan JA, Turkenburg JP, Thomas GH, Hubbard RE, Fischer M., J Am Chem Soc., 2019, 141, 15818-15826 - Statistical thermodynamic foundation for mesoscale aggregation in ternary mixtures
S Shimizu and N Matubayasi, Phys Chem Chem Phys, 2018, 20, 13777-13784. - Hydrotropy and scattering: pre-ouzo as extended near-spinodal region
S Shimizu and N Matubayasi, Phys Chem Chem Phys., 2017, 19,26734-26742 - Unifying hydrotropy under Gibbs phase rule
S Shimizu and N Matubayasi, Phys Chem Chem Phys., 2017, 19,23597-23605 - The origin of cooperative solubilisation by hydrotropes
S Shimizu and N Matubayasi, Phys Chem Chem Phys., 2016, 18,25621-25628 - Hydrotrope accumulation around the drug: The driving force for solubilization and minimum hydrotrope concentration for nicotinamide and urea
J J Booth, M Omar, S Abbott and S Shimizu, Phys Chem Chem Phys., 2015, 17, 8028-8037 - Subnanoscale hydrophobic modulation of salt bridges in aqueous media
S Chen, Y Itoh, T Masuda, S Shimizu, J Zhao, J Ma, S Nakamura, K Okuro, H Noguchi, K Uosaki and T Aida, Science, 2015, 348, 555-559 - Preferential Solvation: Dividing surface vs excess numbers
S Shimizu and N Matubayasi,J Phys Chem B., 2014, 118, 3922-3930. - Hydrotropy: Monomer–micelle equilibrium and minimum hydrotrope concentration
S Shimizu and N Matubayasi, J Phys Chem B., 2014, 118, 10515-10524. - The Mechanism of Hydrophobic Drug Solubilization by Small Molecule Hydrotropes
J J Booth, S Abbott and S Shimizu, J Phys Chem B., 2012, 116, 14915–14921. - Estimating hydration changes upon biomolecular reactions from osmotic stress, high pressure, and preferential hydration experiments
S Shimizu, Proc Natl Acad Sci USA, 2004, 101, 1195-1199.

Contact details
Dr
Seishi
Shimizu
Reader in Biochemistry
York Structural Biology Laboratory
Department of Chemistry
University of York
Heslington
York
YO10 5DD
Tel:
+44 (0)1904 328281
Useful links
