Dr Andrew Holding
Senior Lecturer (Associate Professor) in Biomedical Science
Research
Our research focuses on how cells respond to steroid hormones, both in cancer and in healthy tissues. Through a combination of experimental and computational methods, we find the key parts of the cell that regulate these signals and establish changes that occur between different tissues and in cancer. By understanding these changes, we aim to explain how hormone-driven cancer occurs and why people respond differently to the same treatment.
What are Steroid Hormone Receptors?
Steroids (e.g. oestrogen, cortisol and testosterone) play a vital role both in the growth and development of healthy cells and, when things go wrong, in disease progression. For example, the glucocorticoid receptor plays a key role in the inflammation response; while in two-thirds of breast cancer cases, changes in the oestrogen receptor’s response leads it to become the key driver for tumour growth. Understanding how the cells respond to these hormones, therefore, has huge potential to increase our understanding of disease and to aid the development of future medical interventions.
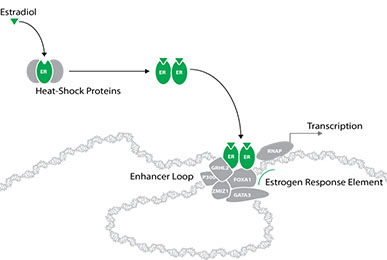
Steroids are detected in the cell by specific steroid hormone receptors, which are found within the cytoplasm and nucleus of the cell. As steroids can diffuse across the cell membrane and into the cell freely, the receptors do not need to be restricted to the outside of the cell membrane. Instead, once the hormone molecule binds to the receptor, the receptor is activated and moves into the nucleus. Here, the receptor binds directly to specific positions on the genomic DNA at specific sequences, called response elements. Once bound, the receptor can then interact with many other proteins to cause the activation and regulation of nearby genes, in turn causing the cell to respond to the steroid.
Previous Research: Elucidating Novel Regulators of the Steroid Hormone Receptors
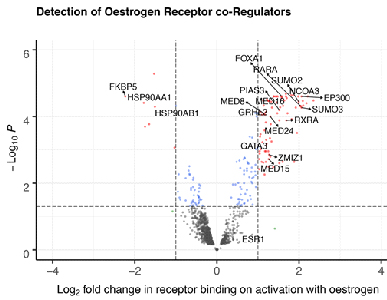
Andrew’s previous research focused on developing methods to identify key proteins that regulate steroid hormone receptors (Holding et al., Genome Biol., 2019 & Papachristou et al., Nat. Comms., 2018) and how the receptors bind to DNA on activation (Guertin et al., NAR, 2018 & Holding et al., eLife, 2018). Using these methods, Andrew has been able to characterise novel proteins that interact with the steroid hormone receptors to regulate their responses. We call these proteins co-regulators.
Current Research: Understanding how these co-Regulators Function Together
Building on these results, our current research utilises CRISPR/Cas9 genome editing combined with single-cell RNA-sequencing to delete the genes that encode for the co-regulators of the steroid hormone receptors and monitor how the loss of these proteins changes the cell’s response to the hormone. By deleting a different gene in each cell, single-cell RNA sequencing enables us to monitor the function of many different co-regulators in a single experiment. We can apply these methods to cells obtained from healthy volunteers and patients. The data we generate are also suitable for machine learning, and these methods enable us to visualise the network of interactions and identify the role of the individual components of the system. We then compare the networks generated in different tissues, individuals or between drug-resistant and sensitive tumours. The results will explain why different cells or individuals respond differently to the same treatment and will provide the groundwork for future therapies that target drug-resistant cancers.