Dr Chris H. Hill
Sir Henry Dale Fellow
Visit Dr Chris H. Hill's profile on the York Research Database to:
- See a full list of publications
- Browse activities and projects
- Explore connections, collaborators, related work and more
Profile
Biography
Chris obtained his PhD in the laboratories of Dr Janet Deane and Professor Randy Read at the Cambridge Institute for Medical Research, studying the structure and mechanism of lysosomal hydrolase enzymes (PNAS 110, 20479-84, 2013; Chem Sci. 6, 3075-3086, 2015; Acta Cryst. F 71, 895-900, 2015). Here, he determined the crystal structure of a tetrameric GALC-SapA complex, defining the molecular basis for glycolipid presentation to hydrolase enzymes by lipid-transfer proteins (Nat Commun. 9, 151, 2018).
In 2015, he joined the MRC Laboratory of Molecular Biology as a Career Development Fellow to work with Dr Lori Passmore on the pre-mRNA cleavage and polyadenylation machinery. By using a combination of X-ray crystallography, cryo-EM and mass spectrometry to study key sub-complexes, he revealed the mechanistic principles of RNA recognition, nuclease activation and inhibition of the host machinery by Influenza A NS1 (Mol Cell 73, 1217-1231, 2019; Science 358, 1056-1059, 2017).
To further explore how viral proteins interfere with cellular information transfer, Chris moved to the Department of Pathology at the University of Cambridge in 2018 to direct a structural and biophysical programme of work for Prof. Ian Brierley on ribosomal frameshifting (Nucleic Acids Res. 47, 8207-8223, 2019). Here, his work on the EMCV 2A protein revealed a novel RNA-binding fold, and a high-resolution cryo-EM structure showed how this protein binds to elongating ribosomes and interferes with translation.
In 2021, Chris was awarded a Sir Henry Dale Fellowship from the Wellcome Trust and Royal Society to establish his own research group at the University of York. Chris is also passionate about undergraduate teaching and is a former Fellow and College Lecturer in Natural Sciences at Queens’ College, Cambridge (2018-2021).
You can find Chris on:
Research
Overview
Recoding in viral infection
Accurate translation of mRNA by the ribosome is essential for all life. It is therefore a high-fidelity process, with spontaneous error rates of only ~ 1 in 100,000 codons. When RNA viruses infect cells, they frequently make proteins that differ from the genetically encoded sequences. These highly-regulated ‘recoding’ events are vitally important to viral gene expression, and if disrupted many viruses (e.g. SARS-CoV-2, HIV-1) fail to complete their replication cycles. A better understanding may therefore present unique opportunities for therapeutic intervention.
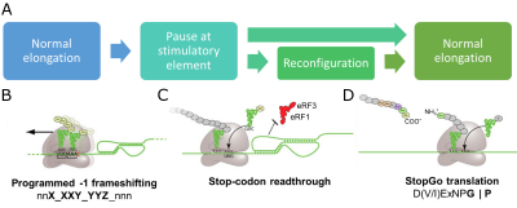
Figure 1. A) Conceptual overview of viral recoding. B-D) Diagrams illustrating key features of PRF, Stop codon readthrough and StopGo translation
All recoding events represent a perturbation of normal elongation, and many involve a pause or stall at a “stimulatory element” before the ribosome resumes normal elongation (Figure 1A). In -1 programmed ribosomal frameshifting (PRF), a proportion of elongating ribosomes shift one nucleotide into the -1 frame, changing the amino acid sequence from that point forward (Figure 1B). During stop codon readthrough, ribosomes fail to recruit release factors at a stop codon, instead extending the protein C-terminally (Figure 1C). In StopGo translation, ribosomes release the upstream peptide and continue translation without re-initiating (Figure 1D). These events are finely-tuned to produce viral proteins in optimal ratios for efficient assembly, and they are regulated by a complex interplay between the elongating ribosome, cis-acting elements in the mRNA or nascent peptide, and trans-acting protein factors.
Our laboratory uses a multidisciplinary approach to better understand recoding, focussing on PRF. We combine classical biochemistry with single-molecule fluorescence microscopy to observe PRF kinetics in real-time both in vitro and in live cells (Figure 2). To reveal mechanistic details, we apply time-resolved cryo-EM, X-ray crystallography, SAXS (Figure 3) and biophysical techniques (Figure 4) to study translating ribosomes, viral proteins, structured RNA elements and RNA-protein interactions at the atomic level (Movie 1).
We are also interested in how cellular proteins act to inhibit recoding events as part of the interferon-stimulated response to viral infection.
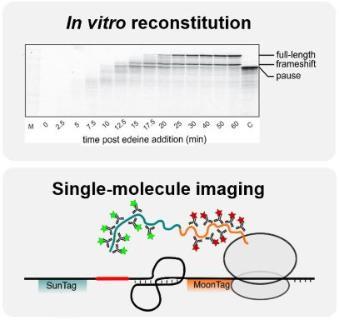
Figure 2. Kinetic measurements of recoding

Figure 3. Structures of ribosomes and stimulatory elements
https://www.youtube.com/watch?v=57dSMT8FbyE
Movie 1. Cryo-EM structure of EMCV 2A protein bound to ribosomes reveals the mechanism of RNA recognition and translational pathology
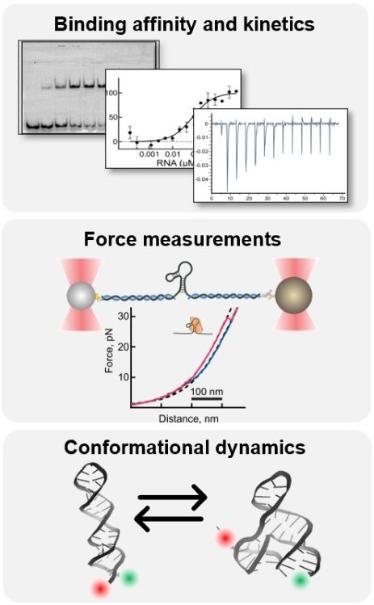
Figure 4. Biochemical and biophysical studies of stimulatory elements
Funding
Wellcome
The Royal Society
BBSRC
Collaborators
Professor Ian Brierley, University of Cambridge, UK
Professor Neva Caliskan, Helmholtz Institute for RNA-based Infection Research (HIRI), Würzburg, Germany
Dr Alex Borodavaka, University of Cambridge, UK


