Dr Gonzalo Blanco
Senior Lecturer
Research
Skeletal muscle main purpose is to produce forces that allow us to fight gravity, move, run or breathe. It is common wisdom that muscles grow or waste depending on the mechanical stresses we place upon them. Muscles make a large proportion of the body and are expensive to maintain, hence it makes physiological sense to maintain proportionality between the amount of tissue and the required force output. However, many gaps exist in our current understanding of how the muscle senses mechanical stress, propagates a signal to the nucleus and elicits transcriptional changes that will eventually adapt its size to the type of activity required.
My research is focused on understanding the mechanisms underlying muscle plasticity. There is no simple experimental approach that could be envisaged to elucidate the many pathways potentially involved in muscle maintenance or hypertrophy. Compared to neurodegenerative diseases, diseases of the muscle are much rarer as any genetic defect that results in a blunted contraction would be lethal. However, rare genetic disorders that compromise muscle plasticity exist and provide an indirect way of identifying key players. The fact that when mutated certain genes cause inherited disorders points to the essential role of the corresponding proteins in muscle maintenance. The subcellular localization of many of these disease-causing proteins has highlighted the Z-disc as one structural component for sensing mechanical stress and modulating muscle performance. Z-discs flank the contractile unit of the muscle fibre, the sarcomere, and are placed perpendicular to the forces applied, an ideal place to detect tension and signal a stress response. KY represents a good example of a Z-disc protein essential for adaptive response of the muscle. The loss of KY protein was shown to cause lack of hypertrophic response to mechanical loads in a mouse model back in 2001. Understanding the molecular mechanisms underlying such lack of plasticity has been a continuous focus in our group. This work paved the way to the identification of recessive mutations in the same gene in humans that are associated with muscle weakness. Moreover, characterizations of the KY protein identified other protein partners such as IGFN1, FLNC or ZAK that also locate to the Z-disc. Some of these proteins have also been shown to underlie very rare myopathic conditions in humans, illustrating how research in mouse models can effectively predict and inform human conditions.
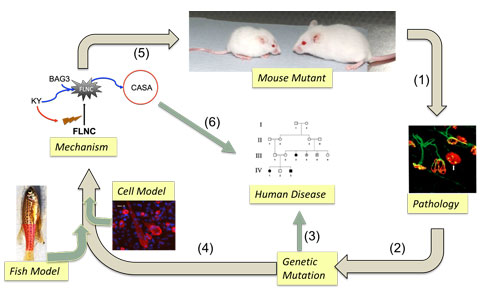
Figure 1. Our experimental paradigm begins with an uncharacterized mouse neuromuscular mutant. The mutation is identified through genetic crosses and next generation sequencing, while detailed histological analyses informs us on the underlying pathology (1 & 2). The causative gene becomes a candidate for inherited human disorders displaying similar pathology of unknown cause (3). Cellular and fish models help us propose a mechanism (4 and 5) that explains the pathological changes observed. Often, these characterizations provide new candidates for human disease (6). In the example shown, the KY protein was first identified in the mouse (Blanco et al., HMG 2001), then congenital human myopathies were identified caused by mutations in the same gene. Recently, fish and cellular models generated by CRISPR/CAS mutagenesis have supported a role for KY in protein turnover of cytoskeletal crosslinkers such as Filamin C (FLNC) (Jokl et al., DMM 2018).
In recent years, mechanisms of protein turnover have been placed at the heart of muscle adaptation. These mechanisms involve the degradation of proteins that get damaged when excessive tension is placed upon them: the higher the loads a muscle support, the more likely structural muscle proteins will irreversibly unfold. Getting rid of damages proteins is essential to prevent formation of aggregates that will otherwise interfere with the contractile machinery of the myofibrils, eventually causing fibre degeneration. One such mechanism is selective autophagy, a cellular process that accumulates specific damaged proteins into digestive vesicles for their degradation. The role of selective autophagy relies on co-chaperones to recognize damaged proteins and direct them towards the degrative pathway. These cochaperones are also found at the Z-disc, supporting the Z-disc role as “tension sensor”.
We have recently shown that KY facilitates the degradation of damaged proteins by selective autophagy using fish, mouse and cellular models. A defective protein turnover is likely to impact the hypertrophic response but other pathways may contribute to this phenotype in KY deficient muscles. A good candidate is the triple kinase ZAK protein, a stress kinase found associated with the KY complex. Stress kinases elicit transcriptional responses to environmental changes through a signaling cascade. Mutations in ZAK have been identified associated with a rare weakening myopathy in humans, indicating the essential role of this signaling kinase in maintaining muscle performance. In collaboration with Betsy Pownall, we are exploring the pathway that ZAK controls in skeletal muscle through the characterization of mouse, fish and cellular models. Our models of KY and ZAK deficiency will contribute to our understanding of the associated human conditions, ultimately helping to develop strategies to tackle these rare diseases.
Teaching and scholarship
![]()
I have spent many years of my professional life in centres of excellence focused on genetics research projects. The experimental paradigm was to use genetic disorders as a window into gene function, exploiting the mouse as model and the neuromuscular systems as the disease target. No matter how common or rare an inherited disorder is, human or mouse genetics tells us that the causative gene must be essential for health. Therefore, imperative to understand well its function. The genetics roulette can be cruel, more so in the case or ultra rare diseases affecting isolated families, as doctors in most situations would have not come across them and are unable to provide effective treatments.
![]()
When I moved to the Department of Biology, a main motivating factor was the opportunity of integrating my research paradigm and results into a new module focused on human genetics. I would not claim it was easy to adapt from talking to specialist audiences. It took continuous effort, student’s feedback and an open mind to develop a solid path that the students could engage with. In my lectures, students learn to interpret complex human pedigrees, calculate genetic risk and learn to apply modern genetics tools to identify the causative mutations using contemporary research papers as the main source. The pace of gene discoveries has increased dramatically with the incorporation of next generation sequencing into the clinic, but many challenges remain. Cracking the mechanism of genetic diseases through interpretation of results from papers is fun, real and engaging. I use examples of my own research to illustrate how the mouse model has been fundamental to ascertain rare genetic diseases in the human population.
![]()
My tutorials are flexible. For new starters, I ask them to come to our first session ready to explain a recent scientific breakthrough in biomedical research as portrait in the media. The group then decides which topic is more appealing and find the papers that led to that discovery. We then critically evaluate them using presentations, summaries and short essays. For Term 2 and onwards, the topic is within the mechanism of muscle plasticity. This may include hypertrophy, atrophy, development and intriguing mechanisms such as “muscle memory”, but core elements in these papers usually overlap with concepts and techniques they come across in lectures such as autophagy, gene editing or mouse conditional knock-outs.
![]()
Projects offered are aligned to my research, generally focusing on a single low risk question and using routine cell biology techniques so that there is a good chance of generating data within the timeframe. Recently, I have also been offering projects that look at the role of arts in science communication. For these, students select the topic they want to communicate, design the media pieces and build a survey to test how effective the process was. These have been unexpectedly popular with students and some have produced outstanding reports.


