Professor Daniela Barillà
Professor of Prokaryotic Genetics
Research
The precise distribution of newly replicated genomes to progeny cells is crucial for stable transmission of genetic information. Bacteria and Archaea employ dedicated DNA segregation factors that drive and deliver low copy number plasmids and chromosomes to specific subcellular locations.
Segregation mechanisms of a multidrug resistance plasmid
One of the model systems under investigation is the multidrug resistance plasmid TP228 that replicates at low copy number in Escherichia coli. This mobile genetic element specifies resistance to a range of antibiotics, including tetracycline, streptomycin and sulphonamides. The plasmid harbours a partition cassette encoding two proteins that are crucial for plasmid inheritance: ParF, a ParA Walker-type ATPase, and ParG, a site-specific DNA-binding protein. We have employed multidisciplinary approaches, spanning from genetics to biochemistry, from super resolution microscopy to biophysics, to investigate the role and interactions of these proteins in mediating plasmid segregation.
|
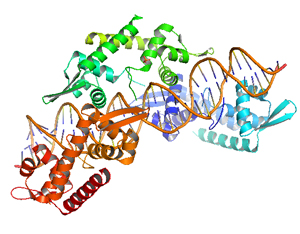
Figure 1. Structure of the ParF protein bound to the ATP analogue AMPPCP (cyan), one monomer is shown in blue-green and the other in red-orange. |
 |
We have solved the three-dimensional structure of ParG and ParF. Crystal structures of the ParF in different nucleotide-bound states suggest a mechanism underlying higher-order structures formation (Fig. 1).
|
Figure 2. A 3D ParF meshwork permeates the nucleoid.3D structural illumination microscopy images of an E. coli cell. ParF is fused to GFP Emerald and the ParG to mCherry. The nucleoid is stained with DAPI. |
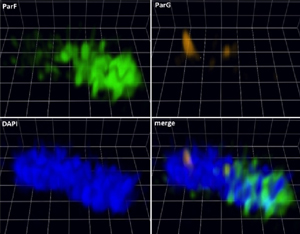 |
Recent microscopy work from our group has shown that ParF assembles into a dynamic three-dimensional meshwork that permeates the nucleoid and periodically shuttles between the nucleoid poles(Fig. 2). We have proposed a Venus flytrap model in which the ParF lattice captures plasmid-ParB analogue complexes within the nucleoid volume and positions them through assembly/disassembly cycles caused by ParF ATP binding and hydrolysis respectively [NAR 2017, 45: 3158-3171].
Discovering the mechanisms whereby multidrug resistance plasmids are inherited will ultimately help us to develop novel therapeutic agents to combat bacterial infections.
Genome segregation in thermophilic Archaea
Archaea are fascinating objects of investigation as they can thrive in very harsh environments thanks to molecular adaptations to a life pushed to extremes. Genome segregation in the archaea domain is virtually an uncharted territory. Two strands of research are currently being pursued, one focusing on mechanisms responsible for chromosome segregation in Sulfolobus solfataricus [Proc Natl Acad Sci USA 2012, 109: 3754-3759] and the other investigating the segregation of a low copy number plasmid in a different Sulfolobus species isolated from acidic hot springs in the island of Hokkaido, Japan. The toolkit for the stable inheritance of this plasmid is a unique three-component machine borrowing building blocks from bacteria and eukarya, including a CenpA-like histone. One of the proteins, AspA, binds to the centromere site, spreading on the DNA and building a superhelix template onto which the other two factors assemble to mediate plasmid segregation [Science 2015, 349: 1120-1124].
|
Figure 1. Structure of the ParF protein bound to the ATP analogue AMPPCP (cyan), one monomer is shown in blue-green and the other in red-orange. |
 |
Teaching and scholarship
![]()
Interacting with young minds is a very rewarding experience. As a teacher, I try to enthuse students and show passion for my subject. I think it is important to prompt them to ‘think big’ and to have the courage of their own ideas.
![]()
As a molecular biologist with a passion for ‘rings’ of DNA, my lectures cover how the chromosome of microbial cells is compacted and delivered to daughter cells upon duplication. Moreover, I teach how genes are turned on and off by proteins acting as repressors or activators. I strive to make my lectures as interactive as possible to engage students by asking questions and then doing straw polls. This approach breaks the rhythm of formal lectures and promotes interaction. I use animations of molecular processes and videos as useful tools to sum up the key messages of lectures. They introduce a ludic element into the lecture: it is teaching and learning without the traditional connotations of teaching. Study questions are included in the last slide of all my lectures and labelled as Mr Quizzer. This is a cartoon-like character who quizzes the students to help them revise concepts covered in the lecture.
![]()
My tutorial topics span from basic biochemistry to genome editing, from synthetic biology to molecular biology of Archaea. These are remarkable objects of investigation due to their exquisitely unusual biological properties and macromolecules. Heat-loving archaea are super microbes thriving at 80ºC and higher temperatures in hot springs, volcanoes, deep sea vents and exhibiting unusual properties, which make these organisms valuable for the development of novel biotechnological applications, but also extremely interesting for studies on life pushed to extremes. During tutorials, students discuss published papers and give presentations, this way honing their communication skills. I also give students the task of organising grant proposal pitches by splitting the group into two teams. This is a really interesting exercise in which the teams assess one another’s plan and provide feedback to the competing team.
![]()
Projects offered to undergraduate and graduate students focus on genome segregation in bacteria and archaea. The approaches involve molecular biology, protein purification, biochemical assays to establish protein-DNA and protein-protein interactions, microscopy, flow cytometry and many other techniques. The nature of the work is molecular and suited to students with an interest in fundamental biological processes.


Contact details
https://barillalabyork.weebly.com/

